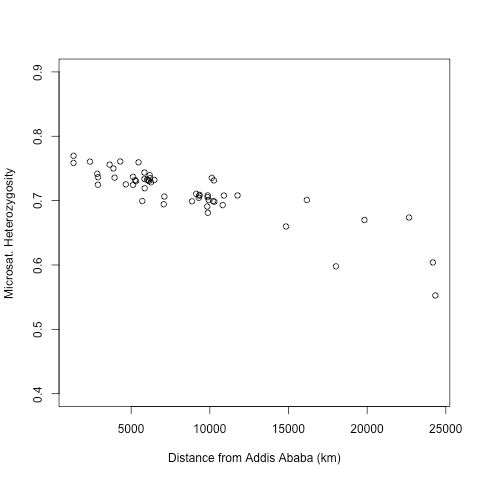
The decreasing level heterozygosity as a function of distance from Africa in human populations is one of the pieces of evidence for the (mainly) out-of-Africa model, see here for a review. This observation is consistent with a serial population bottleneck as populations moved out of Africa and around the world (or in a continuous spatial model increased drift at the front of the wave of advance). Obviously the interpretation of this gradient is complicated by the recent findings of low levels of potential archaic admixture, but the empirical observation still holds and offers an interesting exercise.
In this exercise the students create a serial bottleneck model, and fit its parameters to human data. The exercise gets them to think about modeling the loss in heterozygosity in a different setting, and fit the parameters of a model. The model and data come from Ramachandran et al 2005, thanks to Sohini for sending along the data. If I assign the exercise again I’ll trying and guide the students a little more as they struggled slightly with knowing how to fit the model.
The data file and script are available here.
Exercise:
Download and run the R script: HGDP_He_vs_dist_from_Africa.R. The Human Genome Diversity panel (HGDP) is a set of 53 human populations sampled from around the world, taken from Ramachandran et al. 2005 (thanks to Sohini Ramachandran for sending me these data). The data and graph give the heterozygosity in HGDP populations as a function of the distance from Addis Ababa, Ethiopia. The distances are calculated to avoid large bodies of water. The increasing reduction in heterozygosity has been interpreted as evidence for a set of serial bottlenecks as human populations moved out of East Africa and around the world.
The following model taken from Ramachandran et al. 2005 is a (overly) simple model of human colonization through a series of bottlenecks:
1)An initial population of humans starts in Addis Ababa, with a heterozygosity H.
2) From the Addis Ababa population a new population is founded by D individuals moving a distance R away. This population instantly grows to a very large size (near infinite), with no subsequent migration in or out of the population.
3) From the 2nd population, a 3rd population is founded by D individuals a distance R away, such that this new population is 2R away from Addis Ababa. This 3rd population also instantly grows to a very large size (near infinite size), with no subsequent migration in or out of the population.
This serial founding of new populations continues until a string of populations have been established all the way from Addis Ababa to Brazil (home of the Karitiana people), a distance of over 24000km.
What is the relationship between the heterozygosity and distance from Addis Ababa? Assuming that D is not small, can we hope to distinguish D and R? Using the Ramachandran et al. 2005 data estimate the parameters of this serial bottleneck model. [Useful command: lm() – linear model).
HGDP<-read.table("HGDP_hetsdistfromAfrica.txt",as.is=TRUE,head=TRUE)
plot(HGDP$corrdistEth,HGDP$He,ylim=c(.4,.9),xlab="Distance from Addis Ababa (km)",ylab="Microsat. Heterozygosity")
If you do use these scripts and figures, please acknowledge that fact (mainly so that others can find this resource). Also if you do use them it would be great if you could add a comment to the post so I can see how widely used they are, to get a sense of how worthwhile this is. If you find a bug or make an improved version do let me know.

This work is licensed under a Creative Commons Attribution 3.0 Unported License.