Siblings of the same sex resemble each other to varying degrees. For most traits this is mostly due to differences in the environment between them, and its effects on their development. However, siblings also subtly differ in their genomic similarity, due to the randomness of segregation and recombination. I thought I’d extend our previously discussion of genomic sharing between relatives (see here) to show how variable genomic sharing is between siblings. Again using data from real transmissions.
Below is a picture of the sharing between a pair of sibs. The parent genome is shown as 2 pairs of chromosomes, for each of 22 autosomes. These are coloured by the genomic material they transmitted to the child. The third plot of each row shows the overlap between the siblings’ genomes in light purple. So, for example, the two sibs (on page 1) share all of chromosome 21 as inherited from the father, but only the right tip of the chr21 in the mother. You can also see genomic stretches where the pair of sibs would share their both of their genotype (i.e. both alleles), e.g. the sibs share both maternal and paternal alleles for the first ~1/3 of chr22.
Here’s a slide show illustrating this across 10 pairs of siblings.
I’m posting these as we are currently doing a reading group on recent advances in Quantitative Genetics. This week Gideon and Reid are leading a discussion of Visscher et al “Assumption-free estimation of heritability from genome-wide identity-by-descent sharing between full siblings.”. In that paper they have a plot of the variation of how much of their autosomal genomes siblings share:
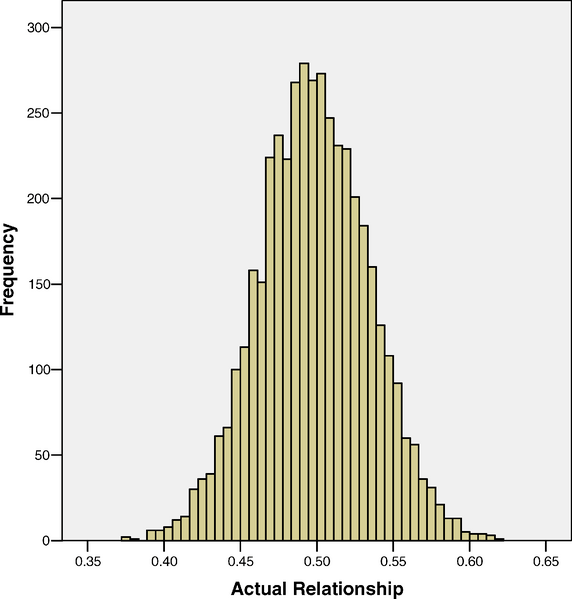
note that the distribution is centered on a half but with a small amount of scatter around that due to the randomness of mendelian segregation and the fact that chromosomes are inherited in big chunks. (I apologize for the default excel graph, but I didn’t make it 😉 )
Visscher et al make really nice use of this slight variability in how much of the genome sibs share to learn about how much variation in height within a population is due to genetic variation. They use the fact that sibs who share slightly more of their genome (>0.5) should have more similar heights, than sibs who share less of their genomes (<0.5). This allows them to partition out how much of the resemblance between siblings is due to a shared environment, as opposed to shared genomes.
This is a really nice application variation in genomic sharing (although the paper is a little tough going in places). It also makes me wonder if sibs are actually unconsciously, weakly aware of these subtle genomic differences (through their similarity in a range of traits, including height etc). I could imagine doing a study where siblings (or others) are asked to assess how similar they are/feel, and then assessing whether this is weakly correlated with the fraction of the genome shared. I keep meaning to followup on this idea with some popgen theory to assess how this might play out in modifying kin-selection and altruism between sibs and other relatives. Anyone know if this has this been looked at before?


somewhat prettier & updated figure in this paper: Vinkhuyzen, Anna AE, et al. “Estimation and Partitioning of Heritability in Human Populations Using Whole Genome Analysis Methods.” Annual review of genetics 47.1 (2013).
Thanks.
Pingback: linkfest – 01/28/14 | hbd* chick