Thanksgiving is over, although you fridge may still be full of leftovers. You probably spent your time wondering exactly what you have in common with your cousin, other than your loathing of brussels sprouts. I’m a British ex-pat so I have no real clue, but I guess that it is what you are pondering as you stare off over the half eaten turkey.
In the previous few posts I talked through the probability you share a given number of genomic blocks with a particular ancestor, and how your number of genetic ancestors compares to your number of genealogical ancestors.
We’ll now take a look at the probability that you and a cousin share a given number of autosomal genomic regions. Every generation you go back the two copies of your genome are spread more and more thinly over your increasing large number of genealogical ancestors. This means that there is a reasonable chance that cousins of more than a few degrees of separation (e.g. 4th cousins, see definitions here) share no autosomal genomic material due to that shared ancestor. This probability of sharing zero increases the further back you and your cousin share a common ancestor.
In the picture below (left panel) is a simulation of the autosomal genome you inherited from your mother (colouring her 2 copies of each chromosome, one from her mother & one from her father). You can see how she transmitted a mosaic of her two copies of each chromosome to you, we call the switches from maternal to paternal chromosome (in your mother) recombination events. In the right two panels I show your genome in your maternal grandmother and grandfather:
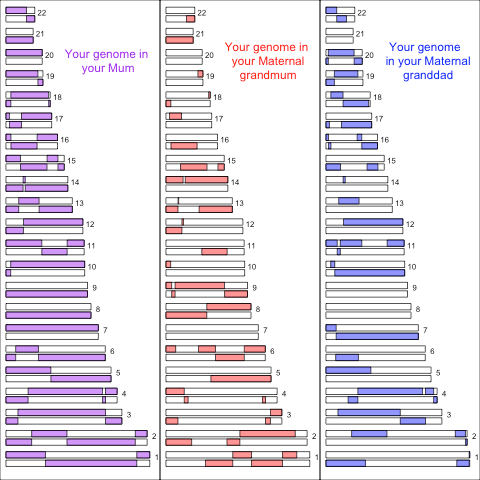
You can see how the genomic chunks they transmitted your mother, and then her to you, are fragmented across their genomes. For example in this simulation your mother passed no genomic material on chromosome 2 on from your maternal granddad, and so all of your maternal chromosome 2 comes from your maternal grandmum.
To illustrate how variable this process is here’s a slide show of 10 other replicates of this process:
The genomic material you share with your first cousin (on your mother’s side) is the overlapping fragments of genome that both of you have inherited from your shared maternal grandparents. In this next plot I show a simulation of you and your cousin’s genomic material that you both inherited from your shared maternal grandparent. In the third panel I show the overlapping genomic regions in purple. (If you are full first cousins you will also have shared genomic regions from your shared grandfather, not shown here.) These are regions where you and your cousin will have matching genomic material, due to having inherited it “identical by descent” from your shared grandmother:

In the cartoon below this is sketched out to show the transmission of the grandparental chromosomes (e.g. chromosome 1) to two cousins, and a stretch of identity by descent (IBD) shared between the cousins is shown:

(Note that the two representations do not show the same outcome of transmission, and so do not match up in terms of the shared genomic material. )
The inherent randomness in the transmission of genetic material, and in where recombination events occur, means that the exact number and location of these shared segments is quite random.
We can also look at more distant cousins. For example, consider second cousins who share a great grandmother. Here’s a simulation of 2nd cousins showing the genomic regions they inherited from their shared great grandmother (following the maternal lineage, mother’s mother’s mother), and the overlap in purple in the final panel:
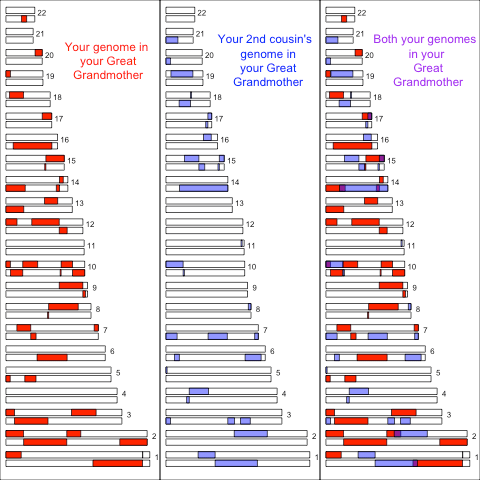
As each of these individual has eight great grandparents, they have inherited less genomic material from their great grandmother than from their grandmother. This material is also broken into shorter blocks as it has been through more generations of recombination. As these individuals inherit less material from their great grandmother there is also less overlapping blocks of “identity by descent” between second cousins than there was between first cousins, and these regions are smaller.
We only have to go back to 4th cousins till it’s quite likely that they share no overlapping autosomal genomic material due to their shared great, great, grandmother. Here’s an example of the material that two fouth cousins inherited from their shared great, great, grandmother:
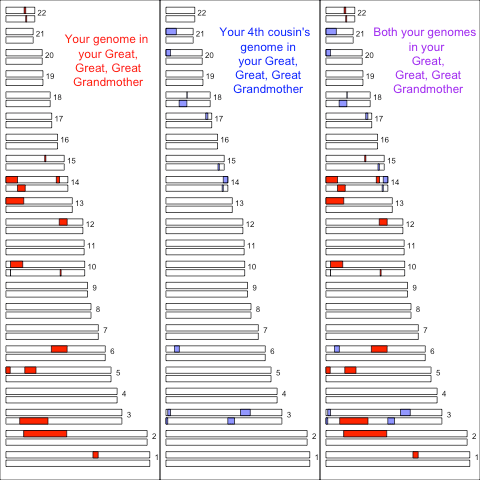
However, by chance they may have some overlapping material inherited from this ancestor. Also you potentially have a reasonably large number of fouth cousins so it is quite likely that you’ll share some genomic material with some of them.
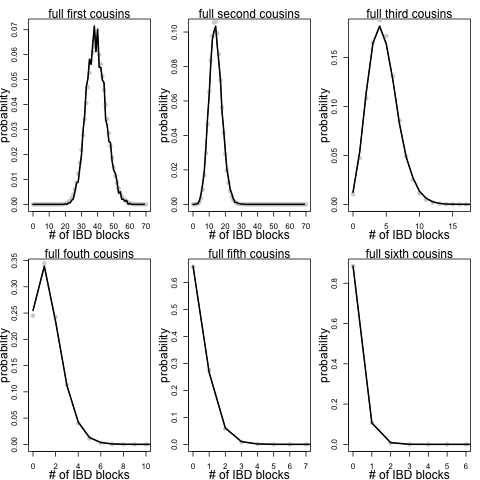
These plots are potentially nice way of illustrating the shared material, but they do not give us a sense of the probability that you share a given number of blocks identical by descent with a cousin. To look at this I simulated a large number of pedigrees and calculated the number of shared autosomal blocks for a variety of depths of relationship for each simulation. Here are the results for full cousins of varying degrees of relationship, the black line shows the results of the simulation (the light grey dots are an analytical approximation that I’ll explain below):
Looking at these we can see the range of numbers of blocks we expect cousins of a given degree of separation to share. For example, roughly 1 in 100 pairs of third full cousins will share zero blocks of their genome due to that shared pair of ancestors. While roughly 25% of pairs of full fourth cousins will share no blocks of their genome due to that pair of shared ancestors. These results assume that this is the only relationship that our cousins share. However, cousins may also share blocks of their genomes identical by descent due to deeper shared relationships (see Peter Ralph and I’s post on this for more discussion of this point). That means for people who share just a couple of blocks, particularly a single block, it may be difficult to assess whether they truly are closely related or whether by chance they have inherited a block from a much more distant ancestor.
We can also make these plots for varying degrees of half sibs:
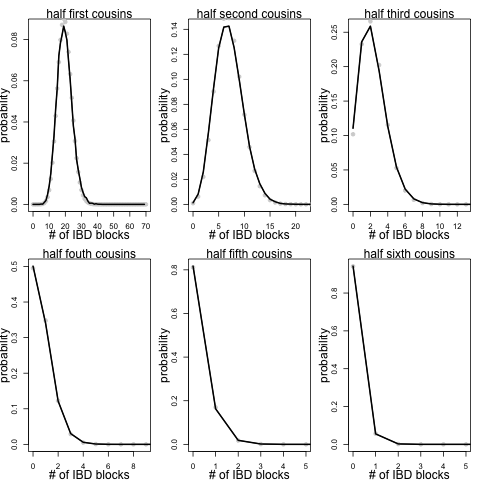
You share likely less with 1/2 cousins of a given degree than you do with full cousins as you only share a single recent common ancestor with these individuals rather than a pair of ancestors.
We can also graph out the probability that you and a relative who share one or two ancestors k generations back share zero blocks:
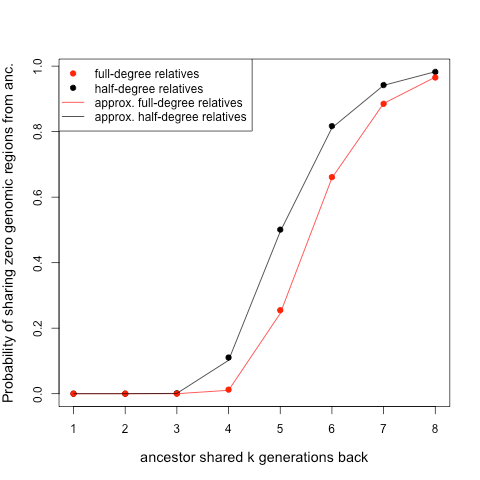
the dots show the simulations, the lines the approximation discussed below.
My simulations are reasonable “exact” (in that they use real recombination data), to get a rigorous analytical answer (albeit ignoring interference) is fairly involved (see Donnelly 1983). However, we can develop a simple, but reasonably accurate, approximation to the expected number of blocks shared between a pair of cousins of a given degree. These approximations have been developed by a number of authors. You can find a reasonable description (open access) in Huff et al, see text above and surrounding equation 7. An elaboration of the ideas laid out here are (I think) used by 23&me and ancestry.com to identify individuals are close relatives in their databases.
We start by considering (say) 1/2 first cousins that share a paternal grandmother but no other recent ancestor. The probability they share a particular block is 1/23=1/8. To understand this probability consider the fact that your grandmother has transmitted one of her two chromosomes in a particular region to your dad, then your dad has to then transmit that region to you (which happens with probability 1/2). For you to share this with your 1/2 cousin, your paternal grandmother also has to transmit the same chromosomal region to your uncle as she did to your dad (that occurs with probability 1/2), and your uncle has to transmit that region to your cousin (probability 1/2 again). Multiplying those probability together we arrive at 1/2*1/2*1/2=1/8. If you were full cousins you would share a particular genomic region with probability 1/4, as you could also share an allele due to your shared grandfather as well as your grandmother (doubling the probability). More generally if you and another individual share a single ancestor d generations ago you share a particular chromosomal region with probability 1/2(2d -1). While if you share two ancestors d generations back (i.e. are full cousins of a given degree) you share a particular chromosomal region with probability 2*(1/22d -1).
That calculation is for a given genomic region, we now have to work out how many different genomic regions you and your cousin could possibly share. You have 22 autosomal chromosomes, and each generation recombination happens in ~34 places on these chromosomes. Looking back d generations your chromosomes are broken up into (22+34d) chunks, which are spread across your ancestors. Likewise your relative’s genome is broken into (22+34*d) chunks. Because recombination events rarely happen in the exactly same place, your two genomes combined are broken into (22+34*d*2) pieces. As each of these is inherited identical by descent to both you and your cousins from that ancestor with probability 1/2(2 d -1), you and your cousins should expect to share 1/2(2 d -1) (22+34d) regions of your genome identical by descent (and double this for full cousins).
A genome does not always undergo ~34 recombination events per generation, this is just the average number. We can approximate the probability distribution of the number of blocks that could possibly be shared between you and a relative by a Poisson distribution with mean (22+68d) as the number of recombination events is roughly Poisson distributed (ignoring recombination interference). As each of these blocks is shared with the probability 1/2(2d -1) for half cousins, the number of shared blocks is Poisson distributed with mean 1/2(2d -1) (22+34d) for half-cousins with an ancestor d generations ago (and double that mean for full cousins). In R we can code up this distribution for 1/2 cousins as dpois(0:70,(33.8*(2*d)+22)/(2^(2*d-1))), where d is the degree of the cousins. This approximation is what is shown as light grey dots in the above figures. This approximation also allows us to get the probability of zero blocks, the lines in the graph just above. For example the probability of zero blocks being shared between two full degree relatives who share two ancestors k generations back is: exp(-2*(33.8*(2*d)+22)/(2^(2*d-1))).
(I’m not totally happy with this description of the approximation, and will think about how to describe it better).

Very cool stuff, as always! However, at the very end, I’m not sure if I follow why the Poisson distribution has mean (22+64d), rather than (22+68d)?
Ahh that’s a typo, apparently I’m not capable of multiplying 34 by 2. Thanks, and glad you like the post.
Graham
Very interesting, Graham. You are acquiring a serious following in the UK.
This is terrific work, thank you.
Now to my specific interest. Norwegian records are very good and very much online and many of us have our lines back to at least the 1500s and the 1300s. What I am finding is more than one large segment of shared DNA with 7th-11th cousins that are doubled and tripled on the same ancestors or two or more distant ancestors. More than 300 years ago perhaps there was less movement of peoples. Still is is very interesting to have so much real data. Eventually I will write it up on my blog!
@Brian Swann
Very interesting, Brian, You ain’t getting a reciprocal serious following among the Geneorata in the U.S. Perhaps, you can go tickle the feet of the POBI people in the UK and get them movin on publishing their study so us Iberian-Welsh-Americans in the U.S. can have more 15th+ cousins in our pedigrees.
Pingback: Papers of the week (w49) | Beyond the Ion Channel
What software are you using to construct your graphics?
They are all done in R, I have my R code available here: https://github.com/cooplab/Genetic_ancestors
This is terrific work, thank you.
In github “https://github.com/cooplab/Genetic_ancestors”, i am not found the file named “recombination_events.out”, So i want to know where can i find it.
Pingback: Episode 207 - DNA Higher Education: CentiMorgans and Segments/ Houses In Photographs
Hi. I think I found this about a year ago but I forgot about who had posted it. I tried using the analytical formula with my known cousins on Ancestry. But for some reason, it worked best if I doubled it. So instead of a factor of 1 for half cousins and 2 for full cousins, I use 2 and 4 respectively. A few from FTDNA and My Heritage seem to follow the pattern. I know it’s just one small sample but I wonder why it works like that. I have one line that’s endogomous, but I excluded them. Maybe my other lines have hidden endogomy. However, the pattern for cM looks normal. So I’ve been using the doubled formula and the table for cM as a check on each other. Any ideas?
Pingback: Episode 207 – DNA Higher Education: CentiMorgans and Segments/ Houses In Photographs
“f you were full cousins you would share a particular genomic region with probability 1/4”: I don’t understand this calculation…the classic coefficient of relatedness among (full first) cousins is 1/8, as in Haldane’s famous quip about laying his life down for 8 cousins. Could you please explain?